We are currently focused on two related and overlapping research projects: Chromatin remodeling factors and transcription coactivators governing enhancer functions in normal development and cancer. We utilize a variety of in vivo and biochemical approaches, involving organismal genetics and cell biology, molecular biology, biochemistry and genomics/bioinformatics. Our genetic model systems include the fruitfly Drosophila melanogaster and mouse hematopoietic stem cells, with projects extending into mouse cancer models and human tumor analyses. We have broad interests in chromatin-based epigenetic gene regulation, signaling pathways in normal and stem cell development and cancer metastases. Training and education of graduate and undergraduate students is a top priority. Our lab is currently funded through grants from the National Science Foundation.
COMPASS complex functions in development and cancer
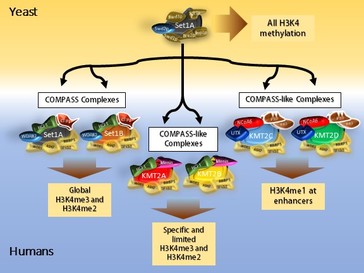
Nuclear receptors, including hormone receptors, activate transcription and reprogram genomes through collaboration with enormous coactivator complexes that recognize, bind and enzymatically modify histone tail residues. These receptors and coactivator complexes (known as COMPASS-like complexes) are critically important regulators of development and differentiation in all higher eucaryotes and disruptions of their functions are strongly linked to developmental defects and many cancers, including lymphoma, medullobastoma (brain tumor), breast, prostate and bladder cancers (see cBio Portal).
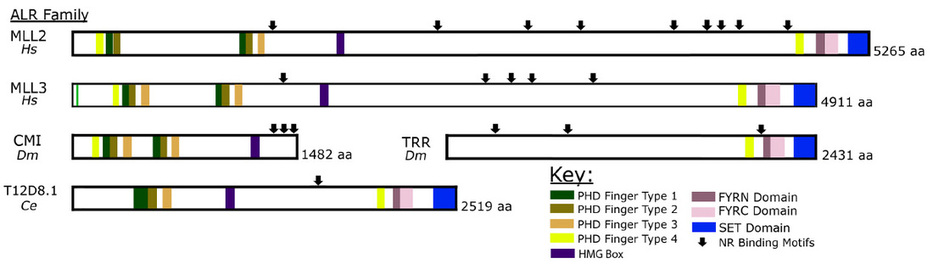
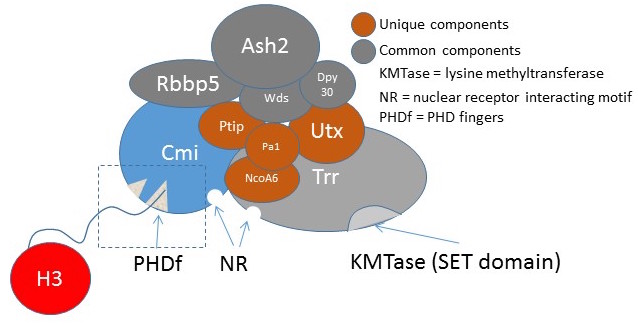
Components of these COMPASS-like coactivator complexes are strongly conserved throughout the animal kingdom, including ALR/MLL2 (KMT2D) and its paralog HALR/MLL3 (KMT2C) that contain both chromatin recognition/binding and histone lysine methyltransferase domains. The genes encoding these proteins arose from a single common ancestor. In Drosophila and some other dipterans, this ancestral gene underwent a fission event, separating into two independent genes, the cara mitad (cmi) gene that we discovered and trithorax-related (trr). The CMI protein represents the N-terminal half of the ALR/MLL2 protein and includes several epigenetic ‘reader’ PHD domains, while the TRR protein represents the C-terminal half and includes the histone methyltransferase activity. This natural split in a genetic model organism has allowed an important opportunity to explore the unique functions of the conserved domains in regulating nuclear receptor functions during development and may allow insights into the roles of nuclear receptor coactivators in human disease.
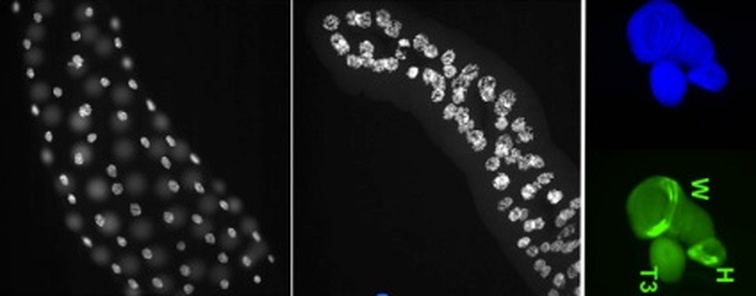
The main research topics of our project include identifying essential functions of cmi in hormone regulated development, and determining how the coordinated functions of CMI and TRR together can read epigenetic histone modifications and further modify those histones during hormone dependent gene activation.
Components of these COMPASS-like coactivator complexes are strongly conserved throughout the animal kingdom, including ALR/MLL2 (KMT2D) and its paralog HALR/MLL3 (KMT2C) that contain both chromatin recognition/binding and histone lysine methyltransferase domains. The genes encoding these proteins arose from a single common ancestor. In Drosophila and some other dipterans, this ancestral gene underwent a fission event, separating into two independent genes, the cara mitad (cmi) gene that we discovered and trithorax-related (trr). The CMI protein represents the N-terminal half of the ALR/MLL2 protein and includes several epigenetic ‘reader’ PHD domains, while the TRR protein represents the C-terminal half and includes the histone methyltransferase activity. This natural split in a genetic model organism has allowed an important opportunity to explore the unique functions of the conserved domains in regulating nuclear receptor functions during development and may allow insights into the roles of nuclear receptor coactivators in human disease.
The main research topics of our project include identifying essential functions of cmi in hormone regulated development, and determining how the coordinated functions of CMI and TRR together can read epigenetic histone modifications and further modify those histones during hormone dependent gene activation.
We are currently focused on comparing the unique and conserved chromatin recognition and binding properties of both the Drosophila and human ALR/MLL family PHD fingers. We examine these questions using genetics, state-of-the-art chromatin binding assays, biophysical measurements and X-ray cystallography/structural biochemistry. We are also collaborating with a group at the University of Chicago to identify all of the genomic targets of the complex so that we can identify how and when the complex activates or blocks gene expression.
Brahma (SWI/SNF) chromatin remodeling complex roles in transcriptional regulation
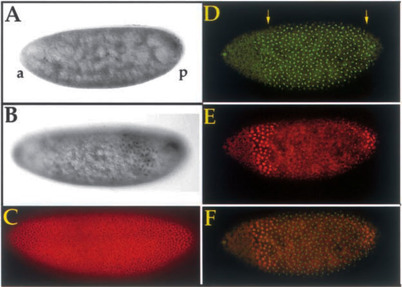
The ability of a cell to access genetic information embedded in the genome is essential for proper development; thus, it is of great importance to understand the cellular processes that guide proper gene regulation in vivo. The nucleosome, composed of a set of core histone proteins and DNA, is the basic building block of chromatin and acts as a primary scaffold for chromosome protection and compaction. Nucleosomes also act as barriers to restrict access to DNA and as a result can serve to prevent the expression of genes and consequently block cell division. Chromatin remodeling is a key step in the process of gene activation through disruptions of DNA-histone contacts that result in changes in DNA wrapping within nucleosomes, referred to as chromatin accessibility. Highly conserved from yeast to vertebrates, chromatin remodeling complexes (such as the SWI/SNF complex) function as essential cofactors to initiate gene transcription and assist in the regulation of transcription elongation as well as alternative splicing that produces different forms of the same gene. Fundamental questions needing to be resolved include how these complexes are targeted to genes in vivo and how they contribute mechanistically to both transcription activation and repression. Although generally acknowledged that chromatin structure can influence how gene transcripts are spliced to produce a variety protein forms from the same gene, many assumptions are made and questions remain concerning the role of the nucleosome in vivo.
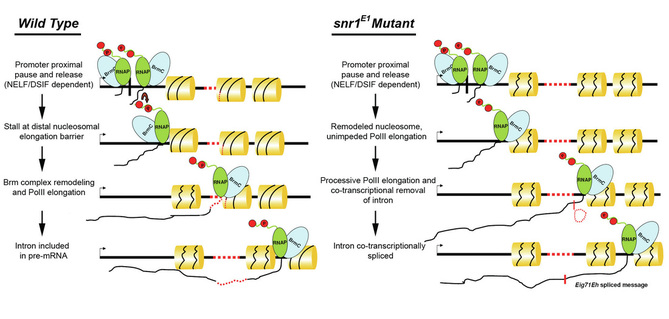
This project tests the hypothesis that chromatin remodeling complexes contribute to gene repression through regulated remodeling of nucleosomes that normally act as impediments to transcription elongation at critical points in development.
This project tests the hypothesis that chromatin remodeling complexes contribute to gene repression through regulated remodeling of nucleosomes that normally act as impediments to transcription elongation at critical points in development.
The research uses two model organisms to address the in vivo functions of nucleosome remodeling complexes using biochemical assays of chromatin compaction, protein binding to chromatin and unique leading edge genetic tools developed for this research study. The main focus of this work is centered on one of the most conserved subunits of the remodeling complex, SNF5, also known as INI1/SNR1/SMARCB1. The Ini1/hSNF5 gene is one of the most potent tumor suppressors ever identified. Loss of the gene has been directly linked to the onset of aggressive childhood malignant cancers of the liver, spleen and pancreas, as well as a large number of pediatric brain tumors. Moreover, approximately 45% of all CD8+ T-cell lymphomas are associated with loss of the Ini1/hSNF5 gene.
Website designed and maintained by Caitlin Dingwall.